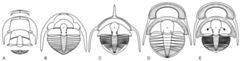
Project 2789: A. Bignon, B. G. Waisfeld, N. E. Vaccari, B. D. Chatterton. 2020. Reassessment of the Order Trinucleida (Trilobita). Journal of Systematic Palaeontology. 18 (13):1061-1077.
Abstract
Trinucleoids have been treated as a superfamily of the Order Asaphida, based on: a pre-occipital glabellar tubercle, a cephalic median suture in most ancestral forms and the presence of broadly similar globular protaspid larvae in the life cycle. Recent discoveries cast doubts on the monophyly of the two last characters. This study provides a phylogenetic analysis whose main purpose is to test the affiliation of trinucleoids. The topology of the strict consensus tree obtained indicates that trinucleoids share a more recent common ancestor with members of the Order Ptychopariida than they do with any asaphid. The stratigraphically higher and most derived trinucleoids are well defined and easy to recognize. Their basal representatives can also be identified with confidence, in spite of substantial variation in the diagnostic characters of the group. These early trinucleoids are similar enough to the typical basal “ptychopariid” bauplan to suggest ancestry within that diffuse and poorly defined trilobite order. Indeed, Ptychopariida is presently regarded as being paraphyletic. We consider that raising trinucleids to ordinal status, by proposing the Order Trinucleida, will make it easier to decipher higher level relationships among trilobites. This action, sustained by improved morphologic and ontogenetic information of Cambro-Ordovician trilobites, reveals convergences of the three characters that have been used to support the inclusion of trinucleids within Asaphida. Based on this new information, we propose six distinct morphotypes of commutavi protaspis and, as a probable sensorial organ, that the cephalic axial tubercle migration might be caused by an increase in the height of the glabella and thus may not bear an important systematic signal. Moreover, the tree topology provided herein offers a preliminary overview of evolution within the order. Alsataspididae and Raphiophoridae are paraphyletic, suggesting that more investigation is needed to define them clearly. Dionidae appears to be more closely related to Raphiophoridae than to Trinucleidae, suggesting that the character state of ‘a perforated fringe’ evolved more than once.Read the article »
Article DOI: 10.1080/14772019.2020.1720324
Project DOI: 10.7934/P2789, http://dx.doi.org/10.7934/P2789
| This project contains | Matrices |
|---|---|
Download Project SDD File | Total scored cells: 4661 Total media associated with cells: 0 Total labels associated with cell media: 0 |
| Characters | |
| Total characters: 75 Total characters with associated media: 14 Total characters with media with labels: 0 Total character states: 185 Total character states with associated media: 130 Total character states with media with labels:0 Total unordered/ordered characters:75/0 |
Currently Viewing:
MorphoBank Project 2789
MorphoBank Project 2789
- Creation Date:
22 November 2017 - Publication Date:
31 December 2019 - Project views: 58527

- Media downloads: 7

- Matrix downloads: 14

Authors' Institutions ![]()
- Universidad Nacional de Cordoba (UNC)
- University of Alberta
- Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET)
- Universidad Nacional de la Rioja
Members
| member name | taxa |
specimens |
media | chars | character
| cell scorings (scored, NPA, "-") | cell
| rules | ||||||
| Arnaud Bignon Project Administrator | 91 | 3 | 151 | 75 | 144 | 0 | 4661 (4276, 0, 385) | 0 | 0 | 0 | ||||
| Brian Chatterton Full membership | 0 | 0 | 0 | 0 | 0 | 0 | 0 (0, 0, 0) | 0 | 0 | 0 | ||||
| Emilio Vaccari Full membership | 0 | 0 | 0 | 0 | 0 | 0 | 0 (0, 0, 0) | 0 | 0 | 0 | ||||
| Beatriz Waisfeld Full membership | 0 | 0 | 0 | 0 | 0 | 0 | 0 (0, 0, 0) | 0 | 0 | 0 | ||||
Taxonomic Overview for Matrix 'M24906' (72 Taxa)
| taxon | unscored cells |
scored cells |
no cell support |
NPA cells |
"-" cells | cell images | labels on cell images |
member access |
| [1] † Myinda uriconii Last Modified in 08/28/19 | 37 | 36 | 36 | 0 | 2 | 0 | 0 | 4 |
| [2] † Myindella crux Last Modified in 08/28/19 | 6 | 67 | 67 | 0 | 2 | 0 | 0 | 4 |
| [3] † Hanchungolithus richthofeni Last Modified in 11/29/18 | 5 | 68 | 68 | 0 | 2 | 0 | 0 | 4 |
| [4] † Ningkianolithus welleri Last Modified in 08/28/19 | 32 | 41 | 41 | 0 | 2 | 0 | 0 | 4 |
| [5] † Reedolithus quebecensis Last Modified in 12/04/18 | 3 | 70 | 70 | 0 | 2 | 0 | 0 | 4 |
| [6] † Protoincaia ancestor Last Modified in 12/04/18 | 10 | 56 | 56 | 0 | 9 | 0 | 0 | 4 |
| [7] † Tretaspis ceryx Last Modified in 12/04/18 | 6 | 60 | 60 | 0 | 9 | 0 | 0 | 4 |
| [8] † Stapeleyella inconstans Last Modified in 12/04/18 | 11 | 55 | 55 | 0 | 9 | 0 | 0 | 4 |
| [9] † Bancroftolithus hughesi Last Modified in 12/04/18 | 0 | 73 | 73 | 0 | 2 | 0 | 0 | 4 |
| [10] † Pragolithus praecedens Last Modified in 11/29/18 | 15 | 51 | 51 | 0 | 9 | 0 | 0 | 4 |
| [11] † Trinucleus fimbriatus Last Modified in 12/04/18 | 6 | 60 | 60 | 0 | 9 | 0 | 0 | 4 |
| [12] † Cryptolithus tesselatus Last Modified in 11/29/18 | 0 | 63 | 63 | 0 | 12 | 0 | 0 | 4 |
| [13] † Deanaspis goldfusii Last Modified in 11/29/18 | 5 | 58 | 58 | 0 | 12 | 0 | 0 | 4 |
| [14] † Onnia s. superba Last Modified in 11/29/18 | 2 | 61 | 61 | 0 | 12 | 0 | 0 | 4 |
| [15] † Marrolithus bureaui Last Modified in 11/29/18 | 2 | 61 | 61 | 0 | 12 | 0 | 0 | 4 |
| [16] † Dionide mareki Last Modified in 11/29/18 | 1 | 62 | 62 | 0 | 12 | 0 | 0 | 4 |
| [17] † Trinucleoides reussi Last Modified in 08/28/19 | 5 | 58 | 58 | 0 | 12 | 0 | 0 | 4 |
| [18] † Ampyxella edgelli Last Modified in 11/29/18 | 9 | 55 | 55 | 0 | 11 | 0 | 0 | 4 |
| [19] † Globampyx trinucleoides Last Modified in 08/28/19 | 3 | 61 | 61 | 0 | 11 | 0 | 0 | 4 |
| [20] † Damghanampyx ginteri Last Modified in 11/29/18 | 6 | 59 | 59 | 0 | 10 | 0 | 0 | 4 |
| [21] † Raphioampyx sinankylosus Last Modified in 12/04/18 | 19 | 45 | 45 | 0 | 11 | 0 | 0 | 4 |
| [22] † Rhombampyx glaber Last Modified in 12/04/18 | 15 | 49 | 49 | 0 | 11 | 0 | 0 | 4 |
| [23] † Raphiophorus parvulus Last Modified in 12/04/19 | 2 | 62 | 62 | 0 | 11 | 0 | 0 | 4 |
| [24] † Cnemidopyge costatus Last Modified in 12/08/18 | 14 | 50 | 50 | 0 | 11 | 0 | 0 | 4 |
| [25] † Lonchodomas tenuis Last Modified in 11/29/18 | 15 | 49 | 49 | 0 | 11 | 0 | 0 | 4 |
| [26] † Ampyx nasutus Last Modified in 08/28/19 | 18 | 46 | 46 | 0 | 11 | 0 | 0 | 4 |
| [27] † Mendolaspis sanjuaninus Last Modified in 11/29/18 | 20 | 44 | 44 | 0 | 11 | 0 | 0 | 4 |
| [28] † Nambeetella fitzroyensis Last Modified in 11/29/18 | 13 | 51 | 51 | 0 | 11 | 0 | 0 | 4 |
| [29] † Pseudampyxina trisegmentata Last Modified in 12/04/18 | 18 | 46 | 46 | 0 | 11 | 0 | 0 | 4 |
| [30] † Lehnertia nawisapa Last Modified in 11/29/18 | 2 | 67 | 67 | 0 | 6 | 0 | 0 | 4 |
| [31] † Pyrimetopus pyrifrons Last Modified in 08/29/19 | 10 | 63 | 63 | 0 | 2 | 0 | 0 | 4 |
| [32] † Orometopus elatifrons Last Modified in 11/29/18 | 13 | 60 | 60 | 0 | 2 | 0 | 0 | 4 |
| [33] † Araiopleura reticulata Last Modified in 11/29/18 | 7 | 66 | 66 | 0 | 2 | 0 | 0 | 4 |
| [34] † Hapalopleura clavata Last Modified in 11/25/17 | 7 | 66 | 66 | 0 | 2 | 0 | 0 | 4 |
| [35] † Seleneceme acuticaudata Last Modified in 12/04/18 | 25 | 41 | 41 | 0 | 9 | 0 | 0 | 4 |
| [36] † Skljarella cracens Last Modified in 12/04/18 | 5 | 68 | 68 | 0 | 2 | 0 | 0 | 4 |
| [37] † Joshuaspis parvus Last Modified in 11/29/18 | 28 | 45 | 45 | 0 | 2 | 0 | 0 | 4 |
| [38] † Doremataspis ornata Last Modified in 08/28/19 | 14 | 59 | 59 | 0 | 2 | 0 | 0 | 4 |
| [39] † Gibbura lepida Last Modified in 08/28/19 | 30 | 43 | 43 | 0 | 2 | 0 | 0 | 4 |
| [40] † Liostracina simesi Last Modified in 11/29/18 | 10 | 63 | 63 | 0 | 2 | 0 | 0 | 4 |
| [41] † Liostracina volens Last Modified in 11/29/18 | 27 | 46 | 46 | 0 | 2 | 0 | 0 | 4 |
| [42] † Liostracina tangwangzhaiensis Last Modified in 11/29/18 | 11 | 62 | 62 | 0 | 2 | 0 | 0 | 4 |
| [43] † Nepea narinosa Last Modified in 08/28/19 | 17 | 56 | 56 | 0 | 2 | 0 | 0 | 4 |
| [44] † Norwoodia boninoi Last Modified in 08/28/19 | 6 | 67 | 67 | 0 | 2 | 0 | 0 | 4 |
| [45] † Glaphyraspis parva Last Modified in 08/28/19 | 14 | 59 | 59 | 0 | 2 | 0 | 0 | 4 |
| [46] † Aphelaspis brachyphasis Last Modified in 11/29/18 | 10 | 63 | 63 | 0 | 2 | 0 | 0 | 4 |
| [47] † Ptychoparia striata Last Modified in 12/04/18 | 3 | 70 | 70 | 0 | 2 | 0 | 0 | 4 |
| [48] † Elrathia kingi Last Modified in 11/29/18 | 5 | 68 | 68 | 0 | 2 | 0 | 0 | 4 |
| [49] † Altikolia kurshabica Last Modified in 08/28/19 | 12 | 61 | 61 | 0 | 2 | 0 | 0 | 4 |
| [50] † Bolaspidella housensis Last Modified in 08/28/19 | 0 | 73 | 73 | 0 | 2 | 0 | 0 | 4 |
| [51] † Kochaspis liliana Last Modified in 08/28/19 | 11 | 62 | 62 | 0 | 2 | 0 | 0 | 4 |
| [52] † Gunnia fava Last Modified in 08/28/19 | 10 | 63 | 63 | 0 | 2 | 0 | 0 | 4 |
| [53] † Modocia whiteleyi Last Modified in 08/28/19 | 4 | 69 | 69 | 0 | 2 | 0 | 0 | 4 |
| [54] † Hardyoides minor Last Modified in 11/29/18 | 7 | 66 | 66 | 0 | 2 | 0 | 0 | 4 |
| [55] † Asaphus (A.) expansus Last Modified in 11/29/18 | 5 | 66 | 66 | 0 | 4 | 0 | 0 | 4 |
| [56] † Megistaspis (m.) limbata Last Modified in 11/29/18 | 14 | 58 | 58 | 0 | 3 | 0 | 0 | 4 |
| [57] † Asaphellus homfrayi Last Modified in 08/28/19 | 7 | 65 | 65 | 0 | 3 | 0 | 0 | 4 |
| [58] † Tsinania canens Last Modified in 08/28/19 | 10 | 62 | 62 | 0 | 3 | 0 | 0 | 4 |
| [59] † Liomegalaspides hupeiensis Last Modified in 11/29/18 | 4 | 65 | 65 | 0 | 6 | 0 | 0 | 4 |
| [60] † Taihungshania shui Last Modified in 12/04/18 | 9 | 60 | 60 | 0 | 6 | 0 | 0 | 4 |
| [61] † Nileus armadillo Last Modified in 08/28/19 | 2 | 68 | 68 | 0 | 5 | 0 | 0 | 4 |
| [62] † Remopleurides xiazhenensis Last Modified in 12/04/18 | 12 | 57 | 57 | 0 | 6 | 0 | 0 | 4 |
| [63] † Pterocephalia norfordi Last Modified in 12/04/18 | 2 | 71 | 71 | 0 | 2 | 0 | 0 | 4 |
| [64] † Maladioides coreanicus Last Modified in 11/29/18 | 10 | 62 | 62 | 0 | 3 | 0 | 0 | 4 |
| [65] † Ivshinaspis reticulata Last Modified in 11/29/18 | 19 | 53 | 53 | 0 | 3 | 0 | 0 | 4 |
| [66] † Haniwa quadrata Last Modified in 11/29/18 | 10 | 62 | 62 | 0 | 3 | 0 | 0 | 4 |
| [67] † Abharella sp. Last Modified in 11/29/18 | 12 | 60 | 60 | 0 | 3 | 0 | 0 | 4 |
| [68] † Ceratopyge forficuloides Last Modified in 11/29/18 | 16 | 56 | 56 | 0 | 3 | 0 | 0 | 4 |
| [69] † Proceratopyge cf. p. lata Last Modified in 11/29/18 | 11 | 61 | 61 | 0 | 3 | 0 | 0 | 4 |
| [70] † Parabolina acanthura Last Modified in 11/29/18 | 4 | 69 | 69 | 0 | 2 | 0 | 0 | 4 |
| [71] † Olenellus gilberti Last Modified in 08/28/19 | 2 | 70 | 70 | 0 | 3 | 0 | 0 | 4 |
| [72] † Redlichia takooensis Last Modified in 12/04/18 | 4 | 68 | 68 | 0 | 3 | 0 | 0 | 4 |
Project views 
| type | number of views | Individual items viewed (where applicable) |
| Total project views | 58527 | |
| Project overview | 1716 | |
| Media views | 43724 | Media search (2638 views); M595224 (294 views); M595225 (299 views); M595202 (349 views); M595246 (264 views); M595195 (368 views); M595206 (348 views); M595177 (355 views); M595264 (283 views); M595277 (266 views); M595245 (269 views); M595180 (373 views); M595194 (363 views); M595239 (263 views); M595223 (332 views); M595350 (262 views); M595214 (330 views); M595199 (347 views); M595184 (372 views); M595269 (288 views); M595175 (353 views); M595272 (274 views); M595248 (263 views); M595203 (354 views); M595268 (289 views); M595278 (294 views); M595196 (345 views); M595192 (356 views); M595229 (266 views); M595226 (301 views); M595227 (278 views); M595276 (260 views); M595249 (282 views); M595251 (281 views); M595218 (312 views); M595182 (359 views); M595270 (283 views); M595244 (269 views); M595198 (329 views); M595179 (361 views); M595235 (273 views); M595260 (281 views); M595193 (347 views); M595242 (280 views); M595222 (331 views); M595238 (283 views); M595253 (285 views); M595262 (290 views); M595185 (359 views); M595230 (283 views); M595261 (289 views); M595266 (280 views); M595197 (339 views); M595237 (268 views); M595220 (317 views); M595219 (329 views); M595212 (329 views); M595240 (264 views); M595232 (267 views); M595252 (275 views); M595234 (269 views); M595250 (263 views); M595241 (281 views); M595228 (283 views); M595274 (287 views); M595191 (357 views); M595279 (281 views); M595221 (341 views); M595265 (283 views); M595201 (361 views); M595205 (333 views); M595187 (357 views); M595207 (353 views); M595233 (307 views); M595217 (318 views); M595204 (341 views); M595213 (351 views); M595211 (352 views); M595208 (342 views); M466122 (360 views); M595189 (351 views); M595243 (292 views); M595176 (378 views); M595178 (360 views); M595183 (354 views); M595190 (358 views); M595200 (310 views); M595209 (345 views); M595210 (370 views); M595215 (339 views); M595216 (336 views); M595231 (286 views); M595236 (298 views); M595247 (268 views); M595259 (254 views); M595263 (263 views); M595267 (268 views); M595271 (266 views); M595273 (262 views); M595275 (278 views); M595351 (261 views); M595352 (194 views); M595353 (196 views); M595354 (196 views); M595355 (193 views); M595356 (199 views); M595357 (200 views); M595358 (197 views); M595378 (202 views); M595379 (199 views); M595380 (202 views); M595381 (202 views); M595382 (170 views); M595383 (193 views); M595384 (198 views); M595385 (203 views); M676252 (201 views); M676253 (213 views); M676254 (195 views); M676255 (195 views); M676256 (203 views); M676257 (205 views); M676258 (199 views); M676259 (200 views); M676260 (205 views); M676261 (196 views); M676262 (196 views); M676263 (200 views); M676264 (180 views); M676265 (201 views); M676266 (203 views); M676267 (200 views); M676268 (201 views); M676269 (206 views); M676270 (194 views); M676271 (197 views); M676272 (193 views); M676273 (193 views); M676274 (194 views); M676275 (190 views); M676276 (212 views); M676277 (201 views); M676278 (203 views); M676279 (192 views); M676280 (197 views); M676281 (207 views); M676282 (195 views); M676283 (171 views); M676284 (196 views); M676285 (200 views); M676286 (197 views); M676287 (191 views); |
| Matrix views | 859 | Trinucleida Matrix (112 views); Matrix landing page (747 views); |
| Documents list | 673 | |
| Bibliography | 579 | |
| Views for media list | 904 | |
| Specimen list | 2110 | |
| Taxon list | 7962 |
Project downloads 
| type | number of downloads | Individual items downloaded (where applicable) |
| Total downloads from project | 165 | |
| Document downloads | 6 | All Trees (4 downloads); Consensus Strict (2 downloads); |
| Project downloads | 138 | |
| Matrix downloads | 14 | Trinucleida Matrix (14 downloads); |
| Media downloads | 7 | M595224 (3 downloads); M595179 (1 download); M595180 (1 download); M595195 (1 download); M466122 (1 download); |